-Search query
-Search result
Showing 1 - 50 of 60 items for (author: saw & je)

EMDB-37963: 
Cryo-EM structure of the hamster prion 23-144 fibril at pH 3.7
Method: helical / : Lee CH, Saw JE, Chen E, Wang CH, Chen R

EMDB-29700: 
Cryo-EM imaging scaffold subunits A and B used to display KRAS G12C complex with GDP
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29713: 
KRAS G12C complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29715: 
KRAS G12C complex with GDP and AMG 510 imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29718: 
Green Fluorescence Protein imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29719: 
KRAS G12V complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO
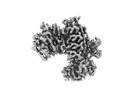
EMDB-29720: 
KRAS G13C complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-28228: 
SARS-CoV-2 spike glycoprotein in complex with the ICO-hu23 neutralizing antibody Fab fragment
Method: single particle / : Yee AW, Morizumi T, Kim K, Kuo A, Ernst OP

PDB-8elj: 
SARS-CoV-2 spike glycoprotein in complex with the ICO-hu23 neutralizing antibody Fab fragment
Method: single particle / : Yee AW, Morizumi T, Kim K, Kuo A, Ernst OP

EMDB-34066: 
F1-ATPase of Acinetobacter baumannii
Method: single particle / : Saw WG, Grueber G

EMDB-15835: 
Cryo-EM structure of homomeric LRRC8C Volume-Regulated Anion Channel
Method: single particle / : Sawicka M, Dutzler R

EMDB-15836: 
Cryo-EM structure of heteromeric LRRC8A/C (1:1 co-transfected) Volume-Regulated Anion Channel in complex with synthetic nanobody Sb1
Method: single particle / : Sawicka M, Dutzler R

EMDB-15837: 
Cryo-EM structure of heteromeric LRRC8A/C Volume-Regulated Anion Channel
Method: single particle / : Sawicka M, Dutzler R

EMDB-15838: 
Cryo-EM structure of heteromeric LRRC8A/C (1:3 co-transfected) Volume-Regulated Anion Channel in complex with synthetic nanobody Sb1
Method: single particle / : Sawicka M, Dutzler R

EMDB-15839: 
Cryo-EM structure of heteromeric LRRC8A/C (endogenous) Volume-Regulated Anion Channel in complex with synthetic nanobody Sb1
Method: single particle / : Sawicka M, Dutzler R

EMDB-15840: 
Cryo-EM structure of homomeric LRRC8A_SAM Volume-Regulated Anion Channel
Method: single particle / : Sawicka M, Dutzler R

EMDB-15841: 
Cryo-EM structure of heteromeric LRRC8A_SAM/C Volume-Regulated Anion Channel
Method: single particle / : Sawicka M, Dutzler R

PDB-8b40: 
Structure of homomeric LRRC8C Volume-Regulated Anion Channel
Method: single particle / : Sawicka M, Dutzler R

PDB-8b41: 
Structure of heteromeric LRRC8A/C (1:1 co-transfected) Volume-Regulated Anion Channel in complex with synthetic nanobody Sb1
Method: single particle / : Sawicka M, Dutzler R

PDB-8b42: 
Structure of heteromeric LRRC8A/C Volume-Regulated Anion Channel.
Method: single particle / : Sawicka M, Dutzler R

PDB-7sxn: 
Orb2A residues 1-9 MYNKFVNFI
Method: electron crystallography / : Bowler JT, Sawaya MR, Boyer DR, Cascio D, Eisenberg DS

PDB-8ddg: 
FYF peptide forms a standard beta-sheet
Method: electron crystallography / : Sawaya MR, Hazari A, Eisenberg DE, Vlahakis NW

EMDB-25026: 
Anaphase Promoting Complex delta APC7
Method: single particle / : Ferguson CJ, Brown NG, Prabu JR, Watson ER, Schulman BA, Bonni A

EMDB-25027: 
Anaphase Promoting Complex delta APC7 + UBE2C,substrate,UbV,CDH1
Method: single particle / : Ferguson CJ, Brown NG, Prabu JR, Watson ER, Schulman BA, Bonni A

EMDB-22738: 
CryoEM reconstruction of SARS-CoV-2 receptor binding domain in complex with the Fab fragment of neutralizing antibody 46
Method: single particle / : Kucharska I, Tan YZ, Benlekbir S, Rubinstein JL, Julien JP

EMDB-22739: 
CryoEM reconstruction of SARS-CoV-2 Spike in complex with the Fab fragment of neutralizing antibody 80.
Method: single particle / : Kucharska I, Tan YZ, Benlekbir S, Rubinstein JL, Julien JP

EMDB-22740: 
CryoEM reconstruction of SARS-CoV-2 Spike in complex with the Fab fragment of neutralizing antibody 298
Method: single particle / : Kucharska I, Tan YZ, Benlekbir S, Rubinstein JL, Julien JP

EMDB-22741: 
CryoEM reconstruction of SARS-CoV-2 Spike in complex with the Fab fragment of neutralizing antibody 324
Method: single particle / : Kucharska I, Tan YZ, Benlekbir S, Rubinstein JL, Julien JP

EMDB-22076: 
Cryo-EM Structure of the Helicobacter pylori dCag3 OMC
Method: single particle / : Sheedlo MJ, Chung JM

EMDB-22081: 
Cryo-EM Structure of the Helicobacter pylori OMC
Method: single particle / : Sheedlo MJ, Chung JM

PDB-6x6k: 
Cryo-EM Structure of the Helicobacter pylori dCag3 OMC
Method: single particle / : Sheedlo MJ, Chung JM, Sawhney N, Durie CL, Cover TL, Ohi MD, Lacy DB

PDB-6x6s: 
Cryo-EM Structure of the Helicobacter pylori OMC
Method: single particle / : Sheedlo MJ, Chung JM, Sawhney N, Durie CL, Cover TL, Ohi MD, Lacy DB

EMDB-22077: 
Cryo-EM Structure of CagX and CagY within the dCag3 Helicobacter pylori PR
Method: single particle / : Sheedlo MJ, Chung JM, Sawhney N, Durie CL, Cover TL, Ohi MD, Lacy DB

PDB-6x6j: 
Cryo-EM Structure of CagX and CagY within the Helicobacter pylori PR
Method: single particle / : Sheedlo MJ, Chung JM, Sawhney N, Durie CL, Cover TL, Ohi MD, Lacy DB

PDB-6x6l: 
Cryo-EM Structure of CagX and CagY within the dCag3 Helicobacter pylori PR
Method: single particle / : Sheedlo MJ, Chung JM, Sawhney N, Durie CL, Cover TL, Ohi MD, Lacy DB

EMDB-21109: 
Proteinase K Determined by MicroED Phased by ARCIMBOLDO_SHREDDER
Method: electron crystallography / : Richards LS, Martynowycz MW, Sawaya MR, Millan C

PDB-6v8r: 
Proteinase K Determined by MicroED Phased by ARCIMBOLDO_SHREDDER
Method: electron crystallography / : Richards LS, Martynowycz MW, Sawaya MR, Millan C

EMDB-0619: 
Amyloid Beta KLVFFAENVGS 16-26 D23N Iowa mutation
Method: electron crystallography / : Griner SL, Sawaya MR

PDB-6o4j: 
Amyloid Beta KLVFFAENVGS 16-26 D23N Iowa mutation
Method: electron crystallography / : Griner SL, Sawaya MR, Rodriguez JA, Cascio D, Gonen T

EMDB-20020: 
Reconstruction of a T4SS OMCC
Method: single particle / : Chung JM, Sheedlo MJ

EMDB-20021: 
PolyAla Model of the PRC from the Type 4 Secretion System of H. pylori
Method: single particle / : Chung JM, Sheedlo MJ

EMDB-20022: 
Reconstruction of a T4SS OMCC
Method: single particle / : Chung JM, Sheedlo MJ, Campbell AM, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

EMDB-20023: 
Reconstruction of a T4SS OMCC
Method: single particle / : Chung JM, Sheedlo MJ, Campbell AM, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

PDB-6odi: 
Structure of CagY from a cryo-EM reconstruction of a T4SS
Method: single particle / : Chung JM, Sheedlo MJ, Campbell A, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

PDB-6odj: 
PolyAla Model of the PRC from the Type 4 Secretion System of H. pylori
Method: single particle / : Chung JM, Sheedlo MJ, Campbell A, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

PDB-6oee: 
Structure of CagT from a cryo-EM reconstruction of a T4SS
Method: single particle / : Chung JM, Sheedlo MJ, Campbell A, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

PDB-6oef: 
PolyAla Model of the O-layer from the Type 4 Secretion System of H. pylori
Method: single particle / : Chung JM, Sheedlo MJ, Campbell A, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

PDB-6oeg: 
Structure of CagX from a cryo-EM reconstruction of a T4SS
Method: single particle / : Chung JM, Sheedlo MJ, Campbell A, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD

PDB-6oeh: 
PolyAla Model of OMCC I-Layer
Method: single particle / : Chung JM, Sheedlo MJ, Campbell A, Sawhney N, Frick-Cheng AE, Lacy DB, Cover TL, Ohi MD
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
